Folding@Home's Exascale Compute Power Helped Unravel Covid's Secrets
Lockdown productivity
Untold millions of people have rushed to render aid to others as the Covid pandemic has gripped the globe, but those individual contributions took different forms. Some of us did what we could and helped by donating spare computing cycles to distributed computing, which helps researchers use the power of everyday home computers and devices to search for cures. Unfortunately, the result of that work isn't always clear in the public eye, but Nature Chemistry has revealed that Folding@Home, the distributed supercomputer to which anyone can lend their system's computing power, enabled researchers to simulate the proteome of the SARS-CoV-2 virus that caused the COVID-19 pandemic.
Folding@Home was introduced in 2000 to allow everyday people to assist with simulations of protein folding to better understand cancer, Alzheimer's disease, and many other medical problems caused by misfolded proteins. So it's been around for decades, but its popularity exploded when the pandemic started.
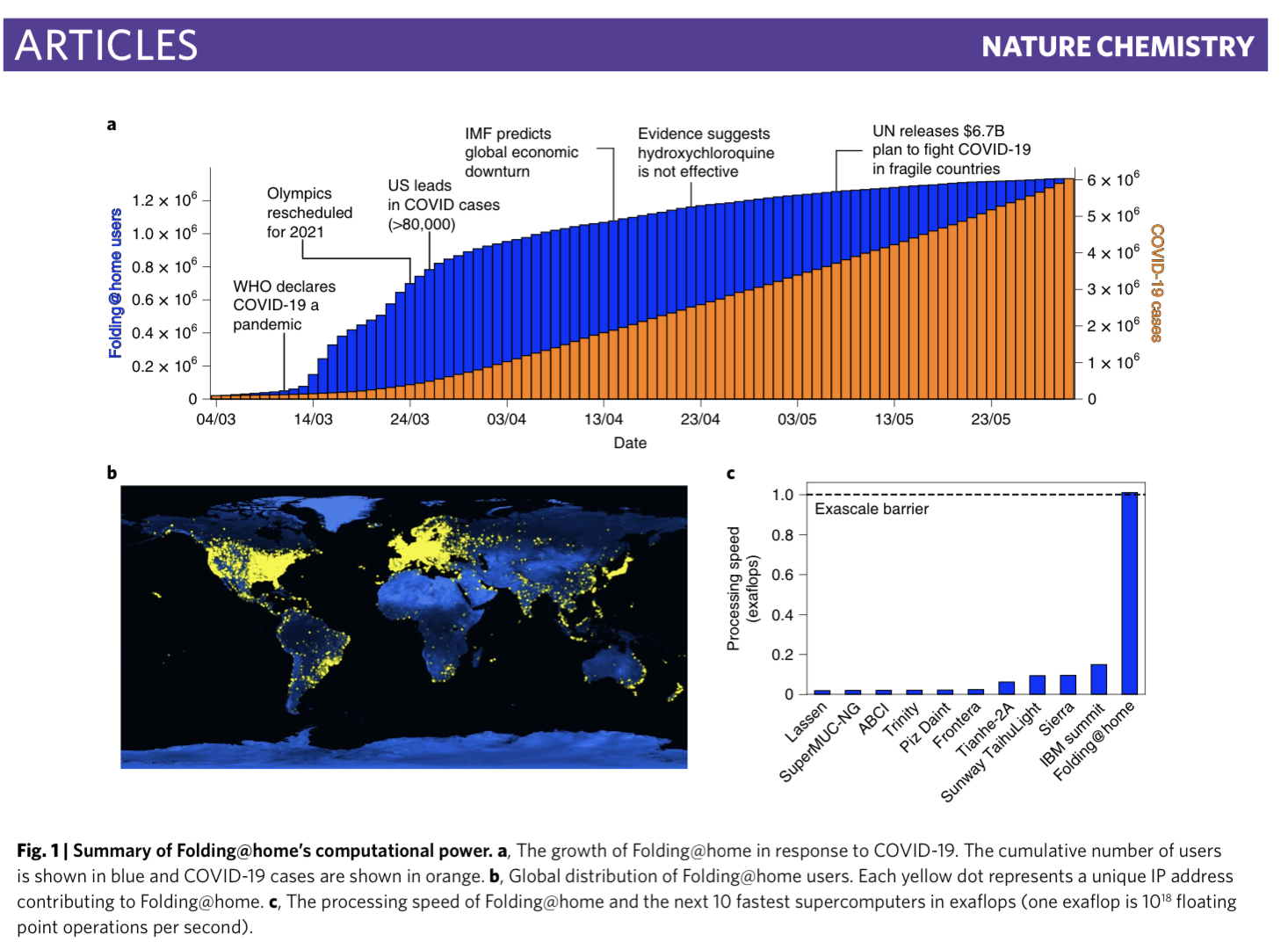
The project's number of active devices grew from 30,000 to 1 million in just three months after COVID-19 went global. Its computing power surpassed that of the top seven supercomputers in March 2020; it broke the exaFLOP barrier a few days later. By April 2020, it was more powerful than the top 500 supercomputers combined.
By then, nobody could deny the power of Folding@Home. The question was how the world's first exascale supercomputer would be used. Performance for its own sake is always intriguing, but much like playing the original Doom with an RTX 3090, it's easy to wonder if that power is being used effectively. Now we have an answer.
"Using this resource, we constructed quantitative maps of the structural ensembles of over two dozen proteins and complex that pertain to SARS-CoV-2 from milliseconds of simulation data generated for each system," researchers said in the Nature Chemistry article. "Together, we have run 0.1s of simulation."
To put that in perspective: An estimated 4.8 million CPU cores and 280,000 GPUs contributed 1.01 exaFLOPs of compute power to simulate SARS-CoV-2, and they were able to simulate a tenth of a second of its proteome. Imagine how long that 0.1 seconds of simulation would've taken on any other platform.
That tenth-of-a-second could also make a significant difference in efforts to combat SARS-CoV-2. The researchers said their revelation of "over 50 'cryptic' pockets" can "expand targeting options for the design of antivirals" and assist with the discovery of "immediate therapeutic interventions" so life can become more normal again.
Get Tom's Hardware's best news and in-depth reviews, straight to your inbox.
The increased power of Folding@Home also has implications beyond the COVID-19 pandemic. "Our work here highlights the incredible utility this compute power has to enable rapid understanding of health and disease," the researchers said, "providing a rich source of structural data for accelerating the design of therapeutics."
"With the continued support of the citizen scientists that have made this work possible," they added, "we have the opportunity to make a profound impact on other global health crises such as cancer, neurodegenerative diseases and antibiotic resistance."
The results of contributing to F@H or other distributed computing networks isn't always as clear-cut as a direct breakthrough that finds a cure, but it does have a tremendous impact — the smaller pieces all build towards the final goal. The work continues. You can learn how to support those efforts on Folding@Home's website.

Nathaniel Mott is a freelance news and features writer for Tom's Hardware US, covering breaking news, security, and the silliest aspects of the tech industry.
-
digitalgriffin F@H is available on multiple platforms, not just windows.Reply
Instead of a mad cash grab built on nothing (crypto), consider helping humanity instead.
During the past year running on a Ryzen 2400g on unraid, I have contributed well over edit 10, 340,000 points work putting me in the top 3% of all contributors. The addition to my power bill was minimal. I worked it out to be about $37/year running 24/7. (Based on increase in power 65 standby watts versus 95 active watts at 12 cents per kWh)
To help humanity, that is a small price to pay. -
epobirs Running F@H on a GPU is far more effective, enough so that I stopped bothering with the CPUs at all. This made keeping the cooling issues under control a lot easier. The down side is my power bill, in the expensive southern CA region, could easily double or triple if I ran all three GPUs among my machines around the clock. Currently that's an RX460 in my main desktop, an RX580 in a less used machine, and an RTX2060 in my gaming system. The RTX2060 is about 3X the effectiveness of the RX580 and about 5X more effective than the RX460. The difference is that the machine with the 460 is the most likely to be on 24/7 for reasons other than folding, and the 460 has a fairly modest power draw, living entirely off bus power with no direct link to the PSU.Reply
I'd run the RTX card more often if I could afford the power bill. The heat also becomes a concern as we head into summer.
As of this moment, this has contributed 110, 543, 803 points to my team (organized by game developer Mark Kern, mostly on Twitter) over a bit more than a year, with a ranking of 17,602 out of 2,875,180 active donors, if I'm reading it correctly. It comes as a bit of a shock to me that I made it so high, considering the far more muscular GPUs out there.
https://stats.foldingathome.org/donor/69377446 -
epobirs Here is a rundown of the recorded performance of GPUs for Folding@home:Reply
https://folding.lar.systems/gpu_ppd/overall_ranks
The link on the same page to CPU performance shows how dramatic the difference is. -
Fade123 Reply
Did you mean 2,000,000,000 points? 2,000,000 points is kinda normal PPD (Points per Day), myself I have contributed with about 245,000,000 points. A friend of mine has contributed insane 49,700,000,000 points!digitalgriffin said:I have contributed well over 2,000,000 points work putting me in the top 3% of all contributors. -
digitalgriffin ReplyFade123 said:Did you mean 2,000,000,000 points? 2,000,000 points is kinda normal PPD (Points per Day), myself I have contributed with about 245,000,000 points. A friend of mine has contributed insane 49,700,000,000 points!
You are right.
10,340,000. But that's just CPU power not the Apu side. There are no iGPU drivers for AMD on unraid. Nvidia yes. AMD no. That's still enough to stick me in the top 3% of contributors in a year's time.
I could run a VM on my pfSense which has a 3400g. But the less I have running on the router the more secure and faster it is. -
RTX 2080 Replydigitalgriffin said:
Instead of a mad cash grab built on nothing (crypto), consider helping humanity instead.
This 100%. I have an RTX 2080 and started contributing 24/7 to folding@home in February of 2020. I actually bought a RTX 2060 KO a few months later so that I could contribute even more. Unfortunately, I had to quit this past January of 2021 (and sell my RTX 2060) due to my super high power bills being a financial strain, but I refuse to use it to mine cryptocurrency like so many people do. It's a stupid waste of resources; imagine the good that could be done if even half the GPUs being used for cryptocurrency were instead running folding@home. -
spongiemaster Reply
The way points are awarded has changed drastically over the years making it far easier to accumulate points today. The poster epobirs above has 110,598,213 pts while completing 3,706 work units. That's a solid contribution. However, the #1 contributor for Tomshardware's team (who hasn't been active for quite a while) has completed 485,465 work units while only totaling 61 million points. Half the points for 131 times the work units completed should give you an idea how much scoring has changed. Looking at the link above for GPU performance, if you have a 3090 and you fold for just one week, you'd almost be in the top 2% (2.16%) for overall points. The point totals are meaningless to the researchers analyzing the results. What really matters is work units completed. Every little bit helps, but folding on a CPU is pretty worthless today compared to the efficiency and speed of GPU's.Fade123 said:Did you mean 2,000,000,000 points? 2,000,000 points is kinda normal PPD (Points per Day), myself I have contributed with about 245,000,000 points. A friend of mine has contributed insane 49,700,000,000 points! -
RTX 2080 Replyspongiemaster said:The way points are awarded has changed drastically over the years making it far easier to accumulate points today. The poster epobirs above has 110,598,213 pts while completing 3,706 work units. That's a solid contribution. However, the #1 contributor for Tomshardware's team (who hasn't been active for quite a while) has completed 485,465 work units while only totaling 61 million points. Half the points for 131 times the work units completed should give you an idea how much scoring has changed. Looking at the link above for GPU performance, if you have a 3090 and you fold for just one week, you'd almost be in the top 2% (2.16%) for overall points. The point totals are meaningless to the researchers analyzing the results. What really matters is work units completed. Every little bit helps, but folding on a CPU is pretty worthless today compared to the efficiency and speed of GPU's.
I understand the point you are trying to make, but actually the reason why modern hardware earns so much more points than hardware in the past did is not because the scoring system has changed, its actually because it has stayed the same.
Allow me to explain. The reason that it is easier to earn more points faster today is because points earned per work unit increase significantly the faster a work unit is finished. If a work unit is completed in one hour instead of two, significantly more than double the points are rewarded for doing it in half the time. The faster a researcher can get a work unit back, the quicker the data can be analyzed; this is why the point system has been set up this way, to incentivise running the fastest hardware as quickly and consistently as possible. Theoretically, the point system could have been set up so that the same amount of points are assigned per completed work unit regardless of how quickly it is finished or that there is a 1:1 progression ratio between time taken to complete a work unit and points rewarded, but instead it was set up to be significantly more rewarding the faster your CPU or GPU can complete a work unit. Thus, the reason powerful modern GPUs can earn the same amount of points in significantly less time than those who have done this for a long time (but on older hardware) is simply a product of the way points are rewarded and have been rewarded all along. -
spongiemaster Reply
Quick return bonuses were not a thing when F@H started. They were introduced 10 years after initial launch. QRB's also have a few requirements before they kick in, including requiring a passkey, returning at least 10 bonus eligible units and returning 80% or more assigned WU's within their time limits. GPU folding has seen significant evolution since its introduction, which again wasn't from day 1. It's only been in recent years that GPU's have basically killed off CPU's as useful folding agents. Nvidia's CUDA driver last year added 15-30% to the PPD for Nvidia GPU's. The folding team will also give extra bonuses to WU's from certain projects deemed higher priority which let's all say it again, has not always been the case.RTX 2080 said:I understand the point you are trying to make, but actually the reason why modern hardware earns so much more points than hardware in the past did is not because the scoring system has changed, its actually because it has stayed the same.
Allow me to explain. The reason that it is easier to earn more points faster today is because points earned per work unit increase significantly the faster a work unit is finished. If a work unit is completed in one hour instead of two, significantly more than double the points are rewarded for doing it in half the time. The faster a researcher can get a work unit back, the quicker the data can be analyzed; this is why the point system has been set up this way, to incentivise running the fastest hardware as quickly and consistently as possible. Theoretically, the point system could have been set up so that the same amount of points are assigned per completed work unit regardless of how quickly it is finished or that there is a 1:1 progression ratio between time taken to complete a work unit and points rewarded, but instead it was set up to be significantly more rewarding the faster your CPU or GPU can complete a work unit. Thus, the reason powerful modern GPUs can earn the same amount of points in significantly less time than those who have done this for a long time (but on older hardware) is simply a product of the way points are rewarded and have been rewarded all along. -
RTX 2080 Replyspongiemaster said:Quick return bonuses were not a thing when F@H started. They were introduced 10 years after initial launch. QRB's also have a few requirements before they kick in, including requiring a passkey, returning at least 10 bonus eligible units and returning 80% or more assigned WU's within their time limits. GPU folding has seen significant evolution since its introduction, which again wasn't from day 1. It's only been in recent years that GPU's have basically killed off CPU's as useful folding agents. Nvidia's CUDA driver last year added 15-30% to the PPD for Nvidia GPU's. The folding team will also give extra bonuses to WU's from certain projects deemed higher priority which let's all say it again, has not always been the case.
That makes sense; I wasn't aware that there have been changes over the years.
Heck, when folding with my RTX 2080, if I get a WU with a base reward of roughly 50,000 points, I'll usually end up getting 200,000 points total upon completion, meaning that my quick return bonus is worth 3x as much as the base amount.
And as you said, GPUs (which can accomplish work units significantly faster than CPUs) have basically killed off CPUs as useful folding agents. Some of the people who have a large number of work units completed compared to points earned made a large amount of their points by folding with CPUs.